You can find data in Pfam in various ways. The Pantoea/Kalamiella strains have the maximum single-nucleotide polymorphisms found within the ISS strains examined. all UniProt and NCBI GI) or different levels of redundancy. Comparative analyses of MAGs and whole-genome sequences of related ISS isolates and their type strains were characterized to understand the variation related to the microbial evolution under microgravity. Pfam full alignments are available from searching a variety of databases, either to provide different accessions (e.g. 
The data presented for each entry is based on the UniProt Reference Proteomes but information on individual UniProtKB sequences can still be found by entering the protein accession. A clan is a collection of Pfam entries which are related by similarity of sequence, structure or profile-HMM. Pfam also generates higher-level groupings of related entries, known as clans. The identification of domains that occur within proteins can therefore provide insights into their function. By contrast, Multiple Sequence Alignment(MSA)is the alignment of three or more biological sequences of similar length. Different combinations of domains give rise to the diverse range of proteins found in nature. Pairwise Sequence Alignmentis used to identify regions of similarity that may indicate functional, structural and/or evolutionary relationships between two biological sequences (protein or nucleic acid). 
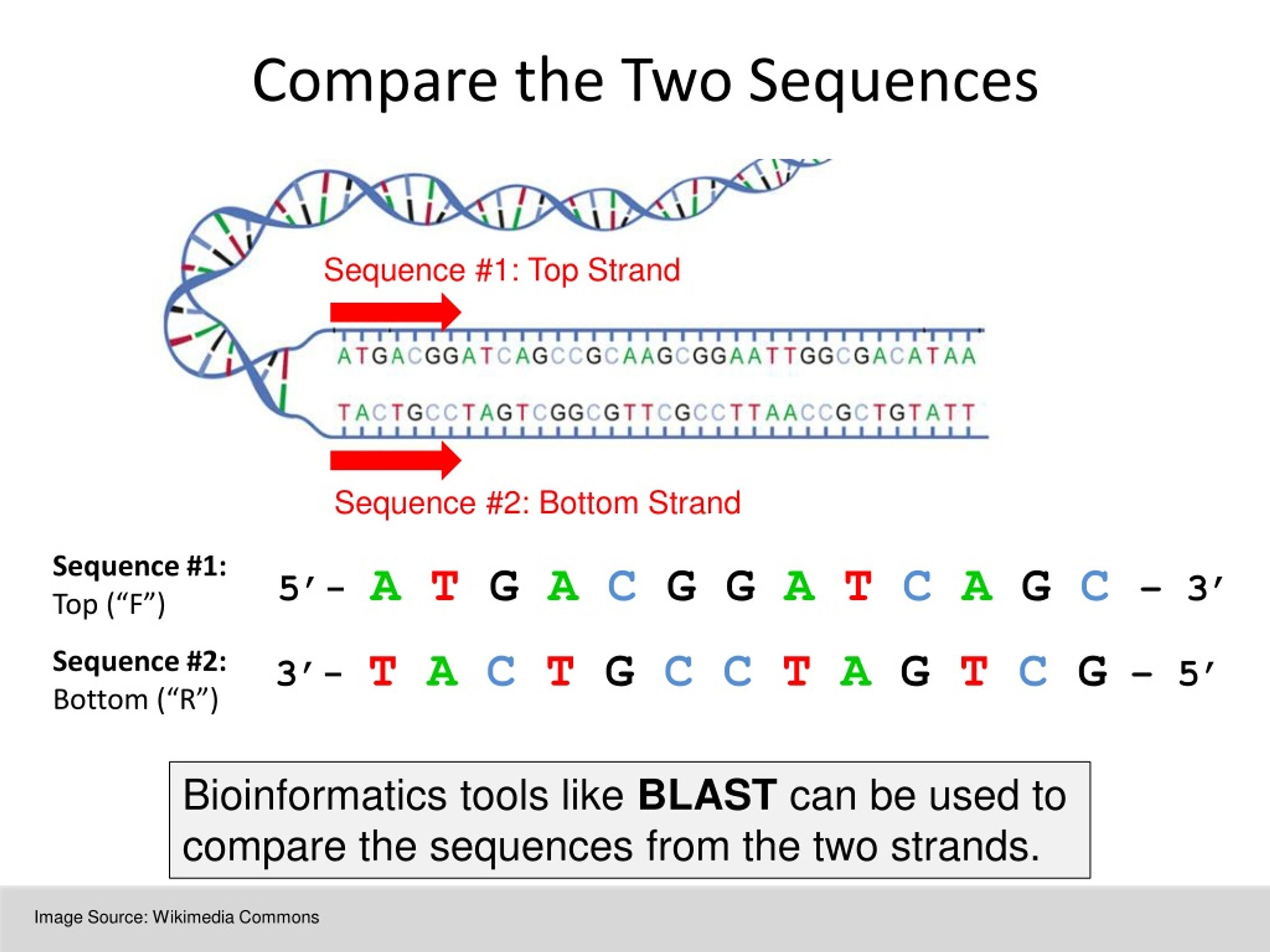
BLAST can be used to infer functional and evolutionary relationships between sequences as well as help identify members of gene families. Proteins are generally composed of one or more functional regions, commonly termed domains. The program compares nucleotide or protein sequences to sequence databases and calculates the statistical significance of matches. The first paper, published in Nucleic Acids Research, introduced the. The data presented for each entry is based on the. Learn more BLAST+ 2.14.0 is here BLASTP, BLASTX, and TBLASTN are faster than before. A clan is a collection of Pfam entries which are related by similarity of sequence, structure or profile-HMM. The program compares nucleotide or protein sequences to sequence databases and calculates the statistical significance. Comparative analysis of DNA sequences from multiple species at varying evolutionary distances is a powerful approach for identifying coding and functional. The Pfam database is a large collection of protein families, each represented by multiple sequence alignments and hidden Markov models (HMMs). MUSCLE (alignment software) MUltiple Sequence Comparison by Log-Expectation ( MUSCLE) is computer software for multiple sequence alignment of protein and nucleotide sequences. Home Recent Results Saved Strategies Help Basic Local Alignment Search Tool BLAST finds regions of similarity between biological sequences.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
